This web page was produced as an assignment for Genetics 564, and undergraduate course at UW-Madison.
Conclusions
Omenn syndrome is a subset of severe combined immunodeficiencies (SCID) characterized by low adaptive immune cell count and autoimmune symptoms. Patients with this disease require a bone marrow transplant soon after birth in order to survive. Additionally, they need to be on immunosuppresants their whole life to manage autoimmune symptoms.
|
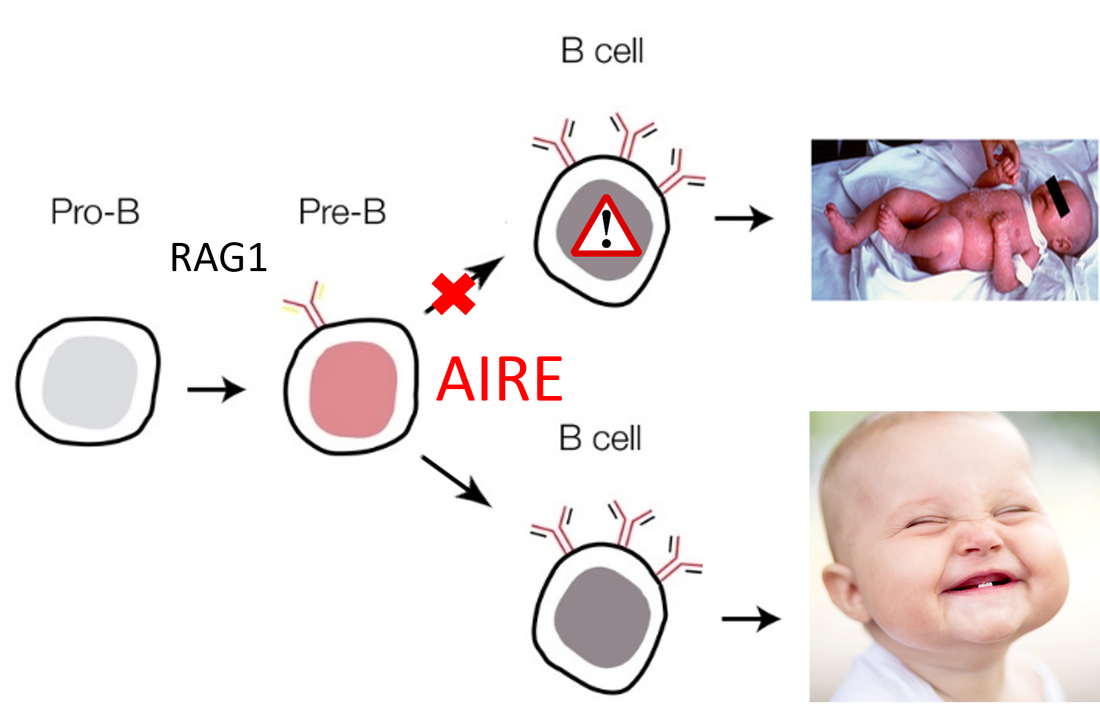
Omenn syndrome is caused by mutations in the RAG1 gene. RAG1 encodes the RAG1 protein, which is involved in T and B cell receptor formation. The protein binds to specific seqeunces in DNA and recombines it to create a diverse library of immune cells that can recognize many different pathogens. Mutations in RAG1 or associated proteins are found in most SCIDs, but does not provide an explanation for autoimmune symptoms. The master regulator of autoimmunity iws the AIRE protein, encoded by the AIRE gene. This gene is a transcription factor that causes the expression of proteins in the thymus and bone marrow that leads to elimination of self-reactive immune cells. The goal of my aims is to explore how RAG1 and AIRE contribute to immune cell development. I hypothesize that mutations that affect pathways involved in immune cell receptor formation and selection are required to result in the Omenn syndrome phenotype.
|
Figure 1: Immune cell development pathway outline
|
AIM 1: To determine changes in gene expression in RAG1 and AIRE double mutants.
My first aim is to demonstrate that mutations in RAG1 and AIRE cause the phenotype found in Omenn syndrome. To test this, I will create mutants strains of mice with the CRISPR-Cas9 system. RAG1 mutations will allow for the leaky immune cell production found in Omenn syndrome. AIRE mutants will have the gene completely knocked out. I will then perform RNA sequencing on each of the mouse lines. We will look at levels of transcripts that have GO terms involved in immune development, B cell selection, T cell selection, and neuronal development. Decreases in proteins involved in immune development will create a phenotype similar to those found typical SCIDS. Decreases in B and T cell selection terms will likely correspond to autoimmune symptoms. Neuronal control will serve as a negative control since these genes are not at all normally associated with this process.
Hypothesis: Mice with mutations in RAG1 and AIRE will have a decreased level of transcripts with GO terms associated with immune development and B cell and T cell selection.
Hypothesis: Mice with mutations in RAG1 and AIRE will have a decreased level of transcripts with GO terms associated with immune development and B cell and T cell selection.
|
Expected results:
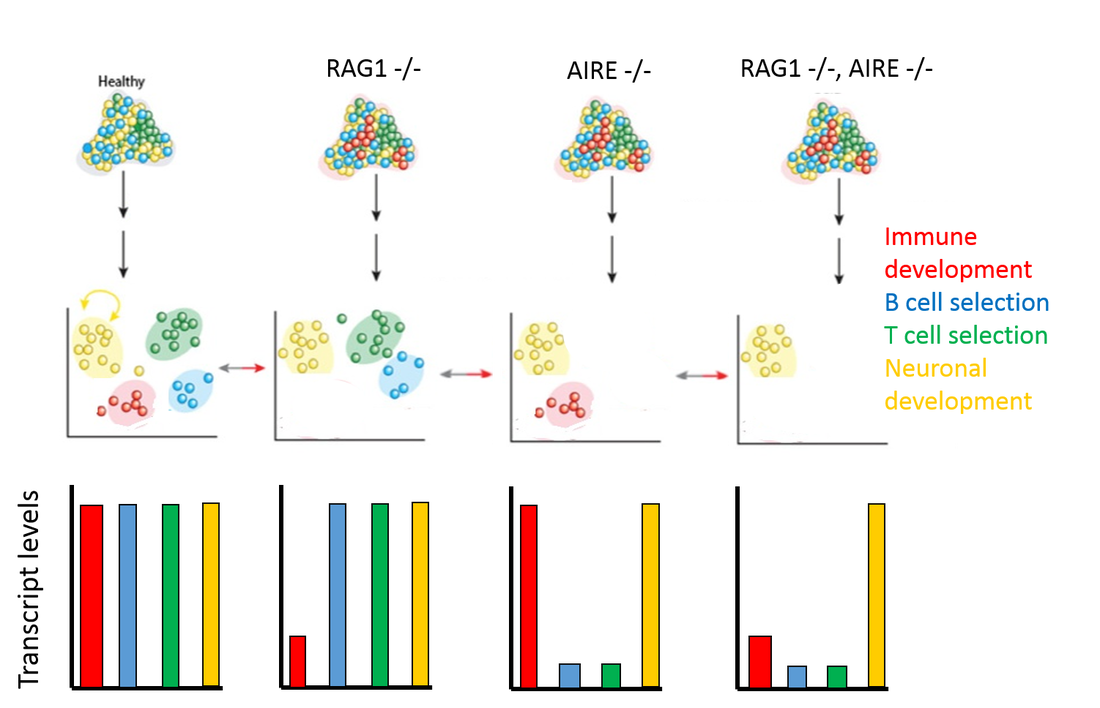
Figure 2 outlines the expected outcome of this experiment. In RAG1 mutants, we expect to see a large decrease in proteins with GO terms involved in immune cell development. For AIRE mutants, there will likely be a large decrease in proteins with GO terms involved in immune cell selection. Our double mutants would likely have a combination of these results, with low amounts of proteins involved in immune selection and development. These mice will also be examined to determine if the phenotype matches with the typical Omenn syndrome phenotype. |
Figure 2: Aim 1 expected results
|
Aim 2: Identify phosphorylation sites in IARE that function in immune cell development.
|
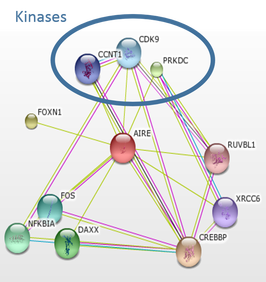
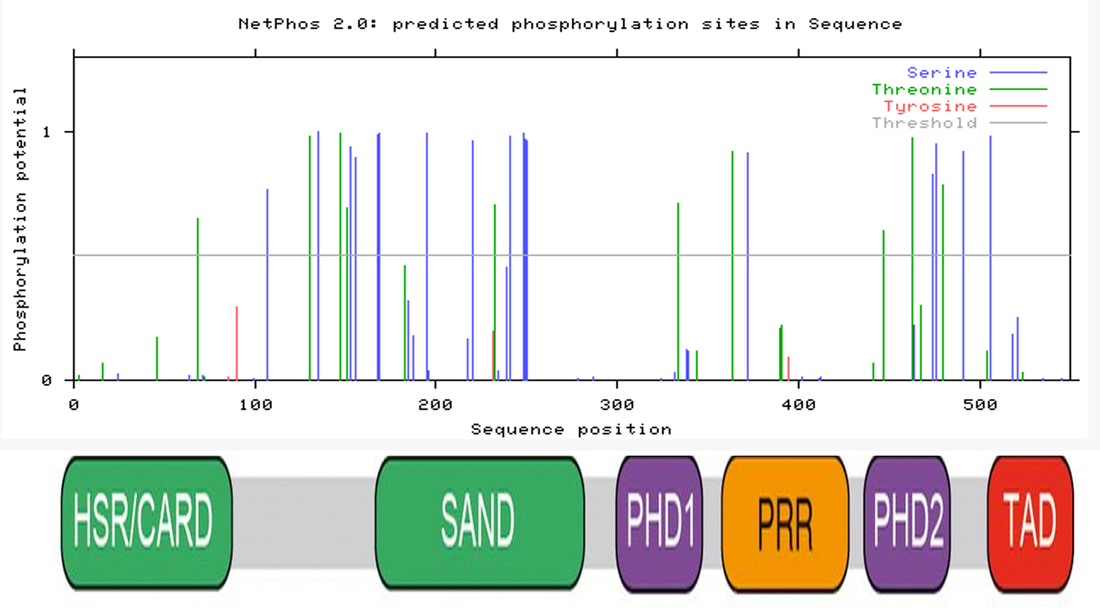
AIRE is involved the master regulator of immune cell selection. I used STRING to generate the protein interaction network for AIRE found in Figure 3. AIRE interacts with several proteins. Interestingly, it interacts with three different kinases. This indicated to me that phosphorylation is likely important in AIRE regulation. I used NetPhos to generate the phosphorylation pattern found in Figure 4. I chose to look at the cluster of phosphorylation sites in the SAND domain, which is involved in DNA binding. There is also a cluster of phosphorylation sites near the PHD2 domain, but this domain is much less conserved than the SAND domain across species, so I eliminated it from my initial evaluation, but this region could be of interest if phosphorylation in the SAND domain does not prove to be essential. To test this, we will design mouse mutants at each of the phosphorylation sites using the CRISPR-Cas9 system and observe the resulting phenotypes.
Hypothesis: Proper phosphorylation of AIRE is necessary for proper protein function. |
Figure 3: AIRE protein interaction network
|
|
Figure 4: Phosphorylation pattern of AIRE as generated by NetPhos
|
Expected results: We expect to find that mutations in the phosphorylation sites of AIRE result in a large increase in autoimmune symptoms. This could mean that a mutation affecting a kinase that normally phosphorylates AIRE could disrupt the entire pathway.
|
Aim 3: Identify shared interacting proteins between AIRE and RAG! important to immune cell development
|
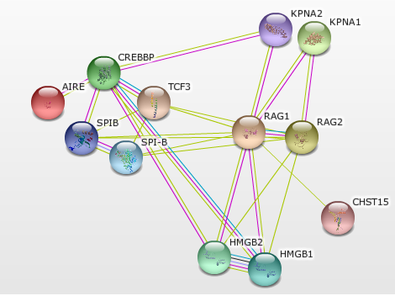
Figure 5 shows a protein interaction network generated by STRING using AIRE and RAG1 as starting proteins. As of now, there are very few interactions that link AIRE and RAG1. If more interactions occur, it could explain how mutations in RAG1 could have an effect on AIRE activity. To identify possible new interacting proteins, a TAP-tag will be used in mice with RAG1 and AIRE as bait. Any proteins that interact with both of the baits will then be assessed using GO terms. Knockout mutants will then be made in mice using the CRISPR-Cas9 system for any proteins that have GO terms related to immune cell development or selection and the phenotypes will be analyzed.
Hypothesis: We hypothesize that mutants in that interact with both RAG1 and AIRE will have abnormal immune phenotypes. |
Figure 5: RAG1 and AIRE protein interaction network
|
Future directions
Patients with Omenn syndrome have a very difficult and often shortened life. The combination of immunodeficiency and autoimmunity creates an obstacle for therapies. Patients receive a bone marrow transplant and then must be on immunosuppressants their whole life. Understanding the mechanism of the disease may be able to aid in treatment for patients with the disease.
Immune cell development as a whole is still a field with a lot of answers. This research could help to clarify exactly how immune cells develop. If protein interactions do prove to be key in signaling between RAG1 and AIRE, it would link immune cell development to selection.
Patients with Omenn syndrome have a very difficult and often shortened life. The combination of immunodeficiency and autoimmunity creates an obstacle for therapies. Patients receive a bone marrow transplant and then must be on immunosuppressants their whole life. Understanding the mechanism of the disease may be able to aid in treatment for patients with the disease.
Immune cell development as a whole is still a field with a lot of answers. This research could help to clarify exactly how immune cells develop. If protein interactions do prove to be key in signaling between RAG1 and AIRE, it would link immune cell development to selection.
| Final talk draft 1 | |
| File Size: | 2484 kb |
| File Type: | pptx |
| final_talk_draft_2.pptx | |
| File Size: | 2461 kb |
| File Type: | pptx |
| final_talk_final_draft.pptx | |
| File Size: | 2290 kb |
| File Type: | pptx |
| final_talk_final_draft.pdf | |
| File Size: | 1179 kb |
| File Type: | |