This web page was produced as an assignment for Genetics 564, and undergraduate course at UW-Madison.
Model Organisms
What makes a good model organism?
A model organism is a well characterized species that is used to model biological systems. Simple organisms, like drosophila melanogaster, can be grown in large numbers, making them great systems to study genetic patterns of inheritance. Zebrafish are a great model organism for studying biological systems because their skin is see through. More complex model organisms like mice, dogs, and apes are good models for studying disease and medicine that can be applied to humans. These organisms provide us a method of performing research in vivo in well known systems [1].
Model organism databases
|
There are a variety of mouse strains that mimic typical SCID phenotypes. These strains are created by adding, deleting, or changing base pairs in the RAG1 gene that renders the gene product useless. These strains can then be manipulated or studied to better understand RAG1 and the SCID phenotype. Recently, Khiong and colleagues accidentally discovered a mutant strain of mice that mimicked the Omenn syndrome phenotype while investigating normal T cell development. This strain could prove to be immensely useful in understanding how Omenn syndrome occurs. [2]
|
There are a variety of other databases for different model organisms. Below are links to several of these.
Discussion
RAG1 is only expressed in organisms with adaptive immune systems. Because of this, many lower level organisms that are popular for modeling diseases and genetic research are not useful for the study of RAG1 and Omenn syndrome. These include drosophila, C. elegans, and popular yeast and plant models. I will look primarily at mice and humans to study the effects of Omenn syndrome.
References
1) Fields S, Johnston M (Mar 2005). Cell biology. Whither model organism research?. Science 307 (5717): 1885–6
2) Marrella, V., Poliani, P.L., Sobacchi, C., Grassi, F., Villa, A. (2008). Of Omenn and mice. Trends in Immunology 29:133-140
2) Marrella, V., Poliani, P.L., Sobacchi, C., Grassi, F., Villa, A. (2008). Of Omenn and mice. Trends in Immunology 29:133-140
Gene expression analysis
Researchers can use mass spectrometry to identify proteins in an individual [1] and Next Generation Sequencing methods to get the genomic sequence of an organism [2]. These are great tools, but at times do not necessarily paint the whole picture of what is happening in a cell. In these cases, researchers can use RNA-sequencing to get an understanding of the genes currently being transcribed, or the transcriptome [3]. There have been a lot of studies that look at the transcriptomes of various organisms under different conditions, known as GEO. Below are the typical results that can be found in this database.
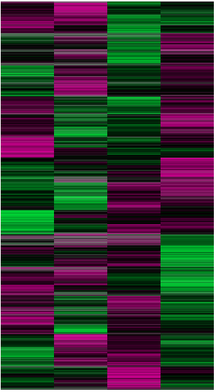
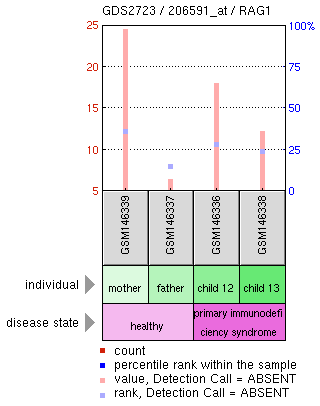
Figure 1: A transcriptomic analysis of of B cells from an individual with a severe combined immunodeficiency. The graph on the left is expression of RAG1 in the B cells. On the right is a heat map of a large number of transcripts from the 4 individuals.
Discussion
As of yet, there has not been a transcriptomic analysis of individuals with Omenn syndrome. I aim to obtain a transcriptome from someone with Omenn syndrome and compare it to the transcriptomes with high risk for autoimmune diseases, SCIDs, and healthy individuals to try to draw conclusions as to the causes of Omenn syndrome.
References
1) Gstaiger, M. and Aebersold, R. (2009). Applying mass spectrometry-based proteomics to genetics, genomics and network biology. Nature Reviews Genetics. 10 (617-627)
2) Metzker, M.L. (2010). Sequencing technologies - the next generation. Nature Reviews Genetics. 11(31-46)
3) Wang, Z., Gerstein, M., and Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics. 10 (57-63)
2) Metzker, M.L. (2010). Sequencing technologies - the next generation. Nature Reviews Genetics. 11(31-46)
3) Wang, Z., Gerstein, M., and Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature Reviews Genetics. 10 (57-63)
Chemical Genetics
Chemical genetics is a novel approach to how we manipulate biological systems. When using something like a mutant strain of a model organism, there are many limitations. Removing the function of a whole protein by deleting it or adding a mutation that renders it useless is an irreversible process. Additionally, proteins tend to be involved in multiple pathways. If a researcher wants to look at how a protein is involved in a single pathway by knocking out its function, they would have to keep in mind that phenotypic differences could be due to its involvement in other pathways. Chemical genetics is a way to avoid these problems. It takes advantage of small molecules that interact with proteins to inactivate them. These small molecules can be targeted to inhibit a single function of the protein. They can also be added later in the life and will in time be removed from the cells. In this way, you can see the phenotype before, during, and after inactivation of a protein. Below are links to the two main databases of small molecules that have been developed for chemical genetics [1].
Discussion
Unforunately, there are no small molecules designed to target RAG1 or AIRE. Once a chemical is synthesized or discovered to inhibit either RAG1 or AIRE, it could provide a very useful tool in studying how deficiencies in these genes affect quantity and specificity of adaptive immune cells.
References
1) Stockwell, B.R. (2004). Exploring biology with small organic molecules. Nature. 432 (846-854)