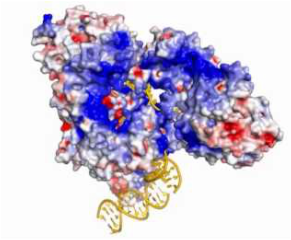
RAG1 protein
This web page was produced as an assignment for Genetics 564, and undergraduate course at UW-Madison.
Domains
What is a domain?
A domain is a conserved part of a gene or protein. These conserved regions are the functional units of a protein. By investigating the domains present in an unknown protein, its function can start to be determined. By looking at the conservation of these domains across the species that have homologous proteins, researchers can learn which parts of the protein are essential and potentially explain small difference between the functions of homologous proteins [1].
Domains in RAG1
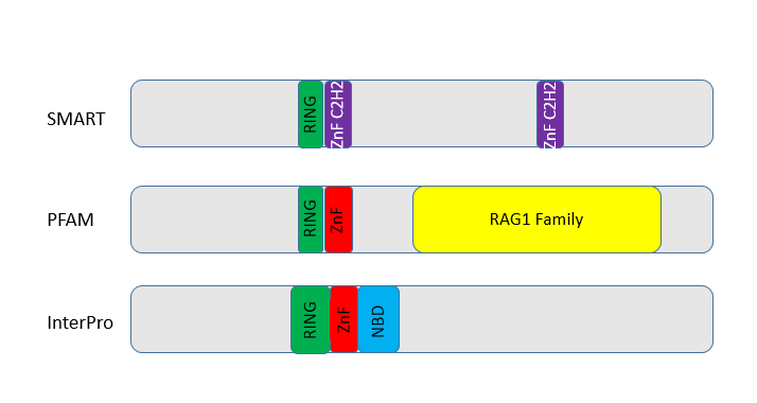
Figure 1: Domains found in RAG1 from three different websites used to find domains.
Figure 1 shows the different domains found in RAG1 using three different programs. The programs use different techniques to find the different domains present in the protein, given an amino acid sequence. Below, you will find a breakdown of what each of the domains means.
RING
A RING domain is a type of Zinc-finger domain that can bind 2 Zinc atoms. Zinc finger domains are short regions that protrude, like fingers, and bind Zinc ions. Many proteins with a RING domain are involved in the ubiquitin pathway. RAG1 uses its RING domain to ubiquinate histones in the DNA and allow it access to the DNA sequences it needs to recombine [3].
ZnF
The ZnF region of RAG1 is another zinc-finger domain. This domain can bind a single zinc atom. This molecule helps it to dimerize to its functional form [2].
RAG1 Family
This a region of the RAG1 gene that PFAM recognizes as being common to RAG1 genes across species [2].
NBD
The nonamer binding domain is the master regulator of RAG1 function. Organisms need prevent recombination from occurring in undesired regions of the DNA as it could lead to mutant proteins that result have undesired consequence, such as cancer. This region of the protein recognizes specific 9 base pair sequences in the genome that, along with 7 base pair sequences recognized by RAG2, mark the regions dedicated for the formation of T and B cell receptors [4].
RING
A RING domain is a type of Zinc-finger domain that can bind 2 Zinc atoms. Zinc finger domains are short regions that protrude, like fingers, and bind Zinc ions. Many proteins with a RING domain are involved in the ubiquitin pathway. RAG1 uses its RING domain to ubiquinate histones in the DNA and allow it access to the DNA sequences it needs to recombine [3].
ZnF
The ZnF region of RAG1 is another zinc-finger domain. This domain can bind a single zinc atom. This molecule helps it to dimerize to its functional form [2].
RAG1 Family
This a region of the RAG1 gene that PFAM recognizes as being common to RAG1 genes across species [2].
NBD
The nonamer binding domain is the master regulator of RAG1 function. Organisms need prevent recombination from occurring in undesired regions of the DNA as it could lead to mutant proteins that result have undesired consequence, such as cancer. This region of the protein recognizes specific 9 base pair sequences in the genome that, along with 7 base pair sequences recognized by RAG2, mark the regions dedicated for the formation of T and B cell receptors [4].
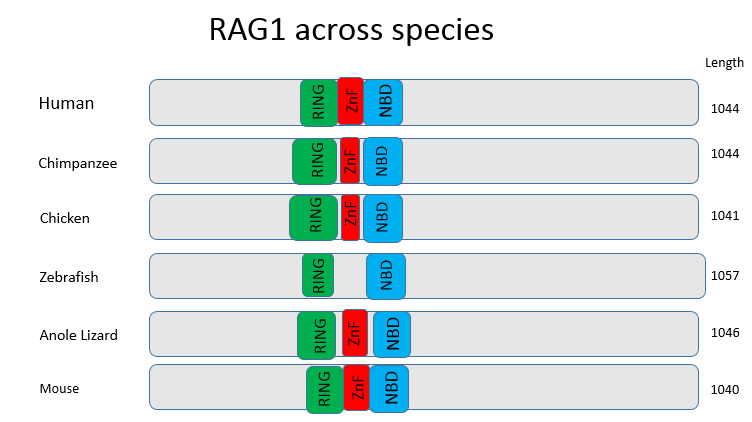
Figure 2: This figure compares the domains found in RAG1 across a variety of organisms. The domains are highly conserved in all species, with the exception of the ZnF domain in zebrafish. These domains were obtained from InterPro, as it was the only domain detecting website that detected the NBD domain.
Discussion
Mutations in RAG1 that lead to Omenn syndrome are located regions encompassing the zinc finger domain and the nonamer binding domain [5]. Mutations in these regions greatly decrease, but do not completely eliminate, the protein's ability to perform these functions. This would result in the "leakiness" of the immunodeficiency found in patients with Omenn Syndrome.
References
1) D'haeseleer, P. (2006). How does DNA sequence motif discovery work?. Nature Biotechnology. 24(8):959-961
2) The Pfam protein families database: R.D. Finn, A. Bateman, J. Clements, P. Coggill, R.Y. Eberhardt, S.R. Eddy, A. Heger, K. Hetherington, L. Holm, J. Mistry, E.L.L. Sonnhammer, J. Tate, M. PuntaNucleic Acids Research (2014) Database Issue 42:D222-D230
3) Grazini U., Zanardi F., Citterio E., Casola S., Goding C.R., McBlane F. (2009) The RING domain of RAG1 ubiquitylates histone H3: a novel activity in chromatin-mediated regulation of V(D)J joining. Mol Cell. 37(2):282-293
4) Yin FF, Bailey S, Innis CA, Ciubotaru M, Kamtekar S, Steitz TA, Schatz DG. (2009). Structure of the RAG1 nonamer binding domain with DNA reveals a dimer that mediates DNA synapsis. Nat Struct Mol Bio. 16(5):499-508
5)Corneo, B., Moshous, D., Güngör, T., Wulffraat, N., Philippet, P., Le Deist, F., Fischer, A., and de Villartay, J.P. (2001).
Identical mutations in RAG1 or RAG2 genes leading to defective V(D)J recombinase activity can cause either T-B–severe combined immune deficiency or Omenn syndrome. Blood 97(9)
Image Reference
RAG1 crystal structure: http://www.nature.com/nature/journal/v518/n7540/full/nature14174.html
2) The Pfam protein families database: R.D. Finn, A. Bateman, J. Clements, P. Coggill, R.Y. Eberhardt, S.R. Eddy, A. Heger, K. Hetherington, L. Holm, J. Mistry, E.L.L. Sonnhammer, J. Tate, M. PuntaNucleic Acids Research (2014) Database Issue 42:D222-D230
3) Grazini U., Zanardi F., Citterio E., Casola S., Goding C.R., McBlane F. (2009) The RING domain of RAG1 ubiquitylates histone H3: a novel activity in chromatin-mediated regulation of V(D)J joining. Mol Cell. 37(2):282-293
4) Yin FF, Bailey S, Innis CA, Ciubotaru M, Kamtekar S, Steitz TA, Schatz DG. (2009). Structure of the RAG1 nonamer binding domain with DNA reveals a dimer that mediates DNA synapsis. Nat Struct Mol Bio. 16(5):499-508
5)
Identical mutations in RAG1 or RAG2 genes leading to defective V(D)J recombinase activity can cause either T-B–severe combined immune deficiency or Omenn syndrome. Blood 97(9)
Image Reference
RAG1 crystal structure: http://www.nature.com/nature/journal/v518/n7540/full/nature14174.html
This web page was produced as an assignment for Genetics 564, and undergraduate course at UW-Madison.